Q1. What are recombinant proteins? How do
bioreactors help in their production?
Solution
The protein produced by genetically altered DNA
(recombinant DNA) in a heterologous host is called recombinant protein. Bioreactors
are vessels in which raw materials are biologically converted into specific products
by microbes. It provides optimum growth conditions such as temperature, pH, substrate,
vitamins, oxygen and salts for the production of recombinant proteins.
Q2. For selection of recombinants, insertional
inactivation of antibiotic marker has been superceded by insertional
inactivation of a marker gene coding for a chromogenic
substrate. Give reasons.
Solution
Selection of recombinants due to
inactivation of antibiotic coding gene is a laborious process as it requires:
(i) a vector with two antibiotic
resistance markers
(ii) preparation of two kinds of media plates,
each with one type of antibiotic.
Transformed cells are first plated on the
antibiotic plate in which the antibiotic gene has not been insertionally inactivated
(example: ampicillin) and incubated overnight for the growth of
transformants. For selection of recombinants, these transformants are
replica-plated on the plate containing second antibiotic plate (example, tetracycline)
which got inactivated due to insertion of the gene). Non-recombinants grow on
both the plates (one carrying ampicillin and the other carrying tetracycline)
while recombinants will grow only on ampicillin plate.
This entire process is laborious and requires
more time (two overnight incubation) as well. However, if we choose
insertional inactivation of a marker that produces colour in presence of a
chromogenic substance, we can easily distinguish between recombinants and non-recombinants
on a single medium plate (containing one antibiotic and the chromogenic
compound) after just overnight growth.
Q3. What is Ti plasmid? Name the
organism where it is found. How does it help in genetic engineering?
Solution
An extra-chromosomal DNA which
delivers gene of interest into a variety of plants and acts as cloning vector
in the host organism is called Ti plasmid. It is present in pathogenic
bacterium Agrobacterium tumefaciens.
Ti plasmid vectors are used for genetic transformation in many dicot plants.
The tumour inducing (Ti) plasmid of Agrobacterium
tumefaciens has been modified into a cloning vector which is no more
pathogenic to the plants, but is still able to use the mechanism to deliver the
gene of interest into a variety of plants.
Q4. How can DNA segments be separated by gel electrophoresis,
visualised and isolated?
Solution
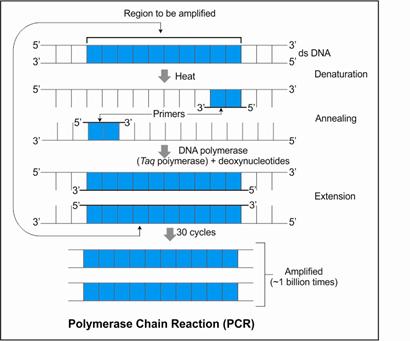
As pure DNA fragments cannot be
seen under visible light, they can be visualised only after staining them
with a solution of ethidium bromide followed by exposure to UV radiation. They
appear as bright orange coloured bands. The separated bands of DNA are then cut
from the agarose gel and extracted by using a convenient technique. This
process is called elution. The eluted DNA fragments are then purified and used
in constructing recombinant DNA by joining them with cloning vectors.
Q5. Explain any two methods of
vectorless gene transfer.
Solution
The
two methods of vectorless gene transfer are as shown below:
(i)
Microinjection: The technique of introducing foreign DNA into a target cell
by injecting the DNA directly into the nucleus with the help of a
micro-needle is called micro-injection.
(ii)
Electroporation: The process in which transient holes are produced in the
plasma membrane of the target cell to incorporate foreign DNA is called
electroporation.
Q6. Who developed the technique of
electrophoresis? State its principle.
Solution
Electrophoresis was developed by
Tiselius in 1937 as a method for the separation of substances with different
ionic properties. It is based on the principle that charged particles move
under the influence of electric current to oppositely charged electrodes.
Q7. How is the amplification of a gene sample of
interest carried out using Polymerase Chain Reaction (PCR)?
Solution
Polymerase Chain Reaction (PCR) is
a technique used to synthesize multiple copies of the desired gene in vitro
within a short span of time.
Amplification of a gene sample of
interest is carried out using PCR in the following three steps:
(a) Denaturation: The double
stranded DNA is denatured by applying high temperature of about 95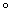 C for 15 seconds. Each separated single stranded
strand now acts as template for the synthesis of new DNA strand.
(b) Annealing: Two sets of primers
are added which anneal at the 3’ end of each separated strand. They mark the
beginning of replication.
(c) Extension: A thermostable DNA
polymerase, Taq polymerase extends
the primers by adding nucleotides complementary to the template provided in
the reaction.
The cycle of denaturation, annealing
and extension is repeated several times to obtain several copies of desired
DNA.
C for 15 seconds. Each separated single stranded
strand now acts as template for the synthesis of new DNA strand.
(b) Annealing: Two sets of primers
are added which anneal at the 3’ end of each separated strand. They mark the
beginning of replication.
(c) Extension: A thermostable DNA
polymerase, Taq polymerase extends
the primers by adding nucleotides complementary to the template provided in
the reaction.
The cycle of denaturation, annealing
and extension is repeated several times to obtain several copies of desired
DNA.

Q8. (i) Mention the number of primers
required in each cycle of polymerase chain reaction (PCR). Write the role of primers
and DNA polymerase in PCR.
(ii) Give the characteristic feature and source
organism of DNA polymerase used in PCR.
Solution
(i) Each cycle of PCR requires two
sets of primers.
Role of primers: Primers are
complementary to the regions of DNA. They anneal to both the strands of DNA
which are then extended using nucleotides.
Role of DNA polymerase: The enzyme
extends the primers using the nucleotides provided in the reaction and the genomic
DNA as template.
(ii) DNA polymerase used in PCR is
thermostable and active even at a temperature of 72 C and is called Taq
polymerase. It is obtained from the bacterium Thermus aquaticus.
C and is called Taq
polymerase. It is obtained from the bacterium Thermus aquaticus.
Q9. What is a cloning vector? Explain the technique
of using such a vector in E.coli.
Solution
Cloning vectors are DNA molecules used
as carriers for transferring a fragment of foreign DNA and capable of replicating
inside the host cell independent of the control of chromosomal DNA. Plasmids and
bacteriophages are commonly used cloning vectors.
The technique of using cloning
vector in E. coli is as follows:
(i) The foreign DNA is linked to
the sequence called ori (origin of replication) and is made to replicate
within the host cell.
(ii) The ligation of foreign DNA is
carried out at a restriction site present in the resistance genes.
(iii) The foreign DNA is then
attached to the plasmid DNA (vector) with the help of enzyme ligase.
(iv) This recombinant DNA (rDNA) is
inserted into the bacterial cell where it multiplies to produce multiple
copies of the desired gene.

Q10. (i) Describe the characteristics that a cloning
vector must possess.
(ii) Why DNA cannot pass through the cell
membrane? Explain. How is a bacterial cell made 'competent' to take up
recombinant DNA from the medium?
Solution
(i) A cloning vector must possess
the following characteristics:
(a) Relaxed origin of replication (ori)
(b) Selectable marker, genes
encoding for an antibiotic resistance or genes encoding for 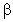 -galactosidase to identify and eliminate non-transformants.
(c) Cloning site or recognition site for the
restriction enzyme to recognise.
(ii) DNA is a hydrophilic molecule. Therefore,
it cannot pass through the cell membrane. The bacterial cells can be made
competent by treating them with a specific concentration of a divalent ion like
calcium. The cells are then incubated on ice followed by a heat shock
treatment by placing them briefly at 42
-galactosidase to identify and eliminate non-transformants.
(c) Cloning site or recognition site for the
restriction enzyme to recognise.
(ii) DNA is a hydrophilic molecule. Therefore,
it cannot pass through the cell membrane. The bacterial cells can be made
competent by treating them with a specific concentration of a divalent ion like
calcium. The cells are then incubated on ice followed by a heat shock
treatment by placing them briefly at 42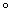 C and then putting back on ice.
C and then putting back on ice.
Comments
Post a Comment